- Press Office
- Press releases 2020
- The secret life of microbes – a snapshot of molecules in a deep-sea symbiosis
The secret life of microbes – a snapshot of molecules in a deep-sea symbiosis
Bacteria in our environment can be difficult to study: They are tiny and often live under conditions hard to recreate in the lab, for example in the deep sea or as symbionts in an animal host (or both, as the symbiotic bacteria in the present study). Investigations of the bacterial genome tell us what the microbes are theoretically capable of. What they actually do, however, is not revealed. Thus, scientists study the so-called metabolome of the bacteria: It comprises every metabolite the cells produce or consume, for example proteins, sugars or fats.
A team of researchers around Benedikt Geier and Manuel Liebeke from the Max Planck Institute for Marine Microbiology in Bremen has now developed a method to identify individual bacteria and at the same time determine which metabolites are present in the cells. With the new method they investigate how bacteria live and survive as symbiotic tenants in deep-sea mussels. Liebeke and his group analysed hundreds of metabolic products on an area smaller than one square millimeter. This enables them to understand how the symbiotic microbes live and communicate in their host. “We virtually take a snapshot of bacteria at work – just as it functions in its natural environment, here within a single animal cell,” says Liebeke. “And we can do this with an impressive resolution of a few micrometers, about ten times thinner than a human hair.”
Snap-frozen for the best snapshot: Not just what is happening, but also who is involved!
„For our analyses, we use mussel tissue that has been snap-frozen and can thus be cut into wafer-thin slices,“ Benedikt Geier explains. „From these slices, we take a snapshot of the chemical compounds of the cells using a special mass spectrometry technique called MALDI-MS imaging. When analysing this snapshot in detail, we are able to distinguish many different metabolites on a very small area.“ They provide information about which metabolites the bacteria use for what purpose and how they cohabit with their mussel host. In addition to the MALDI-MSI at the Max Planck Institute in Bremen, Liebeke and his team used a new MS imaging prototype at the Justus Liebig University in Gießen in close cooperation with Professor Spengler, which enabled particularly high-resolution insights.
Correct conclusions from the images of the metabolites are only possible if we also know who produces or uses them. “To date, we have only been able to measure the metabolites," explains Geier, "but we did not know whether any bacteria were involved and if so, which ones.” To solve this problem, the researchers added a second technique, the so-called fluorescence in situ hybridization, or FISH, to identify individual bacterial cells in the same sample. “The combination with FISH was the key for us to interpret the high-resolution MALDI-MS images in a meaningful way and correlate them with the bacteria in the mussel tissue.”
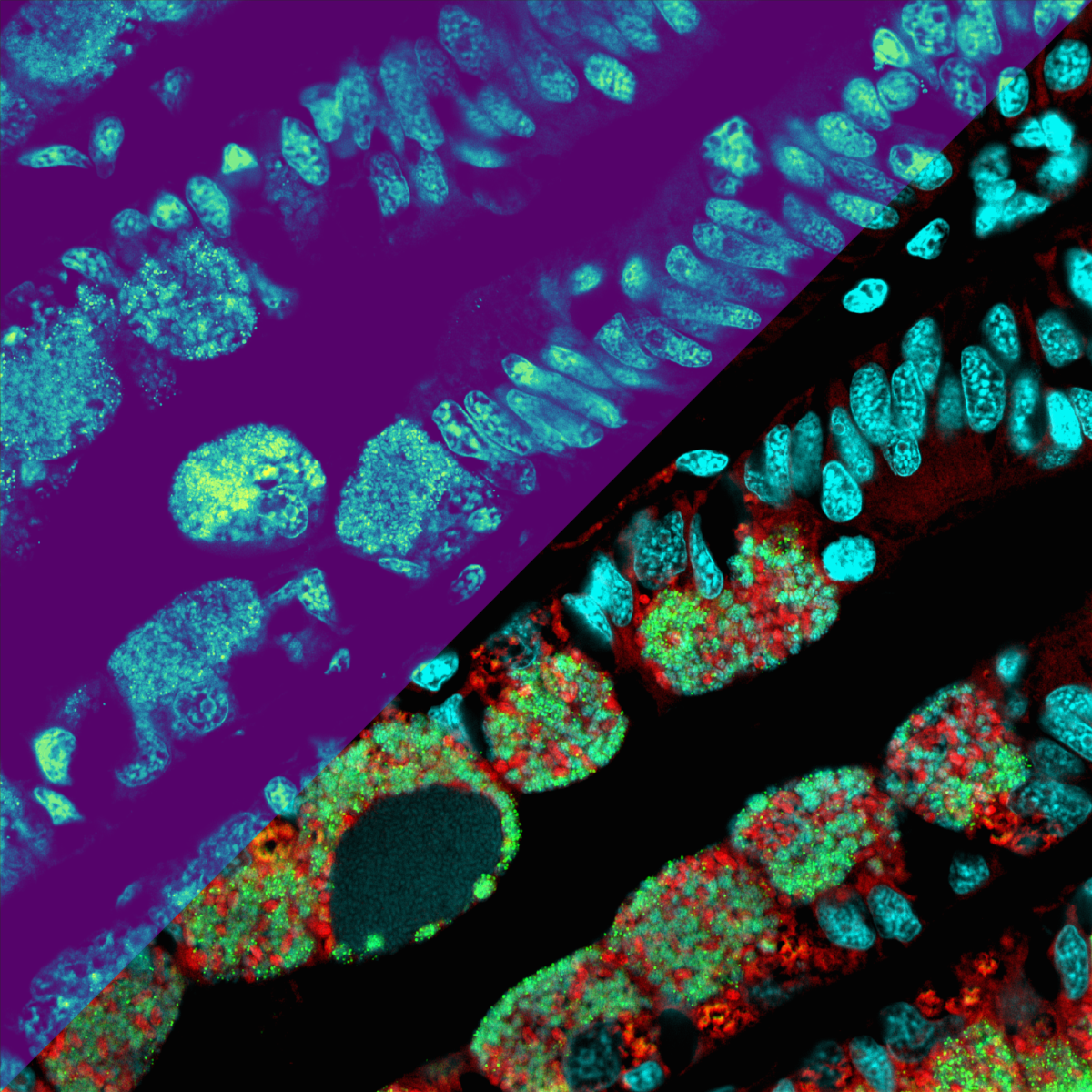
From the deep sea, now on camera
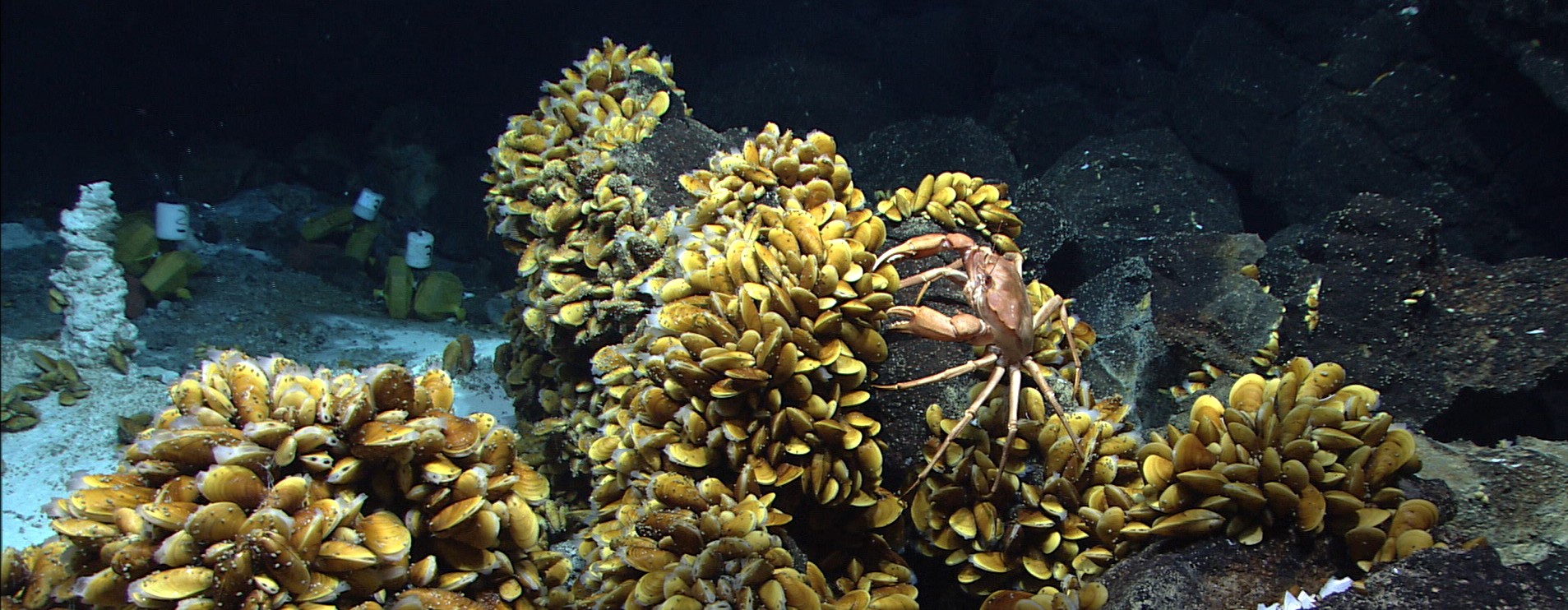
For the present study, Geier and his colleagues used samples from black smokers in the deep sea – towering chimneys where hot, mineral-rich water gushes out of the seafloor. Animals and bacteria can only survive there in symbiotic community. Geier investigates the coexistence of bacteria and mussels, in particular the close linkage of their metabolism, as part of his PhD-thesis. With the new method, he was able to show that the composition of lipids in the mussel differs significantly in body regions with and without bacterial tenants. “Up to now, we were not able to gain such insights as the samples were homogenised, that is virtually blended, before analysis,“ Geier explains. “Moreover the fact that our method works on samples coming directly from the environment and not from the lab underlines its great potential,“ he continues.
All plants and animals as well as us humans live in association with microorganisms, sharing metabolites through close interactions. “Applying this method in other host-microbe interactions will allow for many exciting new insights into the secret life of microbes. I am curious to see whether we can use this imaging approach for looking into the shared chemistry between microbes and organs in whole animals. There is still a lot to discover!”
Further reading
Award winning methodology
For this innovative method, Benedikt Geier received the MSI Award, which is presented annually for outstanding scientific work that has been achieved using mass spectrometry imaging (MSI) techniques. You can read more about this here.
Original publication
Benedikt K. Geier, Emilia Sogin, Dolma Michellod, Moritz Janda, Mario Kompauer, Bernhard Spengler, Nicole Dubilier, Manuel Liebeke: Spatial metabolomics of in situ host–microbe interactions at the micrometre scale. Nature Microbiology.
Participating institutions
- Max Planck Institute for Marine Microbiology, Bremen, Germany
- MARUM, Center for Marine Environmental Sciences, University of Bremen, Germany
- Justus Liebig University Gießen, Germany
Please direct your queries to ...
Group leader
MPI for Marine Microbiology
Celsiusstr. 1
D-28359 Bremen
Germany
|
Room: |
3244 |
|
Phone: |

Head of Press & Communications
MPI for Marine Microbiology
Celsiusstr. 1
D-28359 Bremen
Germany
|
Room: |
1345 |
|
Phone: |


