- Departments
- Department of Symbiosis
- Manuel Liebeke
Dr. Manuel Liebeke
Group leader
MPI for Marine Microbiology
Celsiusstr. 1
D-28359 Bremen
Germany
|
Room: |
3244 |
|
Phone: |

My Main Research Interests
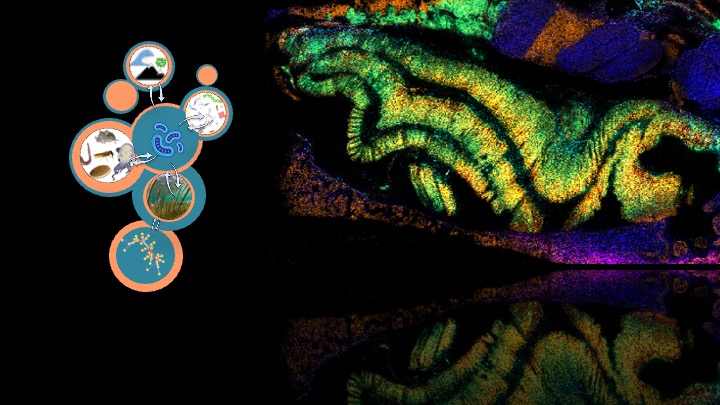
Metabolites serve as a mirror of active metabolic pathways and offer a direct window into the physiological state and ecological roles of microorganisms. The‚Metabolic Interactions‘ lab is at the forefront of microbial metabolomics research, using mass spectrometry to uncover novel insights into microbial communities across different scales. We develop key methods for marine metabolomics involving high-resolution mass-spectrometry imaging with microscopy to reveal the spatial metabolome and community structure of microbial systems, providing a deeper understanding of microbial-host co-metabolism and the specific roles of metabolites in different ecological contexts. Highlighted in our recent finding that sugars accumulate in the rhizosphere of seagrasses through metabolic interactions with plant phenolics. Ultimately, our vision is to transfer spatial metabolite data from single cells into multicellular and environmental contexts, leading to a more comprehensive understanding of microbial ecosystems.
My group develops high-resolution Spatial Imaging Methods for in situ measurements of metabolites in host-microbe systems named spatial metabolomics.
Our major approaches are:
- spatial metabolomics on host-microbe systems
- Mass spectrometry Imaging on the µm scale
- correlative imaging approaches,
- e.g. combining MALDI-MSI with FISH to map bacteria
- 3D scanning approaches like µCT to get 3D anatomy of hosts
- environmental metabolomics
- marine metabolomics, measure metabolites in seawater
- tissue extracts of animals, plants and microorganisms directly sampled in the field
- sediment porewater metabolites
- stable isotope tracer experiments
- following the fate of 13C or deuterated metabolites in symbiotic systems with MS